Guy Hanke
“I know of nothing sublime that is not some modification of power” Edmund Burke


Research Philosophy
Background
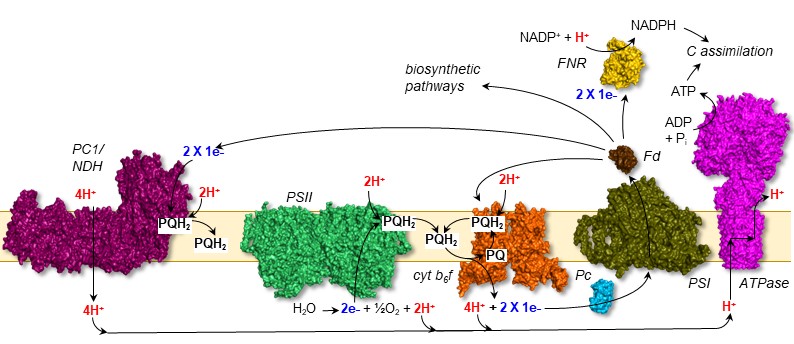
Figure 1. Photosynthetic electron transport. Light excitation of photosystem II (PSII) and photosystem I (PSI) results in electron transport through protein complexes in the thylakoid membrane. This drives proton pumping, resulting in a gradient that can be harvested for the synthesis of ATP. Ferredoxin (Fd) and Fd:NADP(H) oxidoreductase are at the pivotal point in either distributing electrons to metabolic pahways or returning them to the chain in a cyclic electron flow.